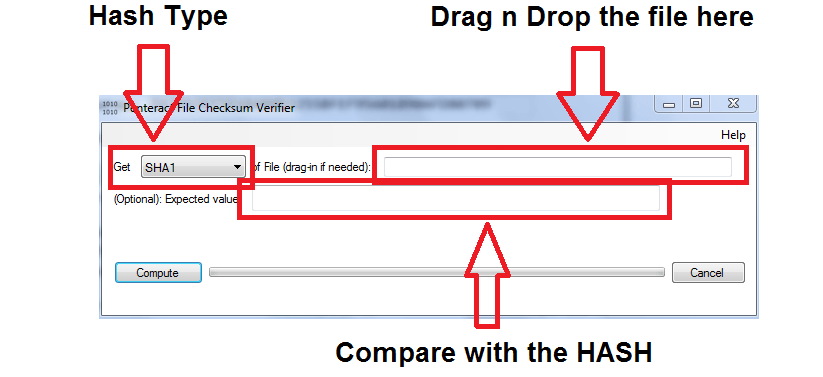
If necessary for the use case, the “relative mode” by executing e.g. The recipe assumes that the absolute paths of the files are the same on source as well as on target system. Windows users may use the included PowerShell function “Get-FileHash”
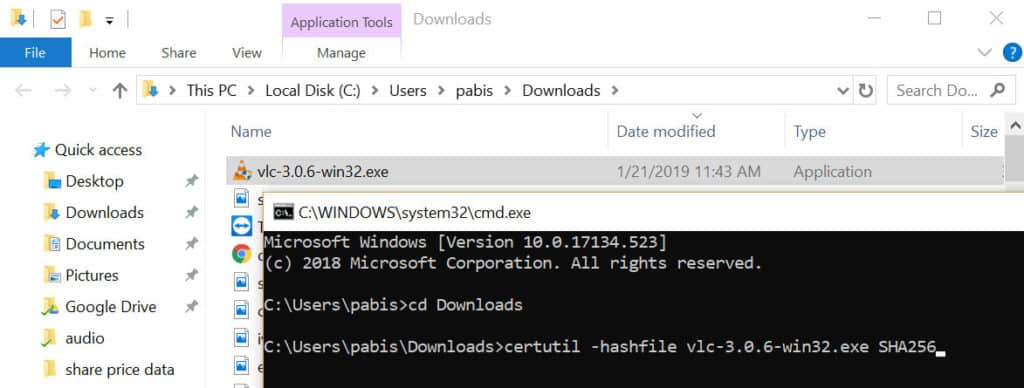
(hint: the Linux tool has the -tag flag which generates macOS-compatible output.) Their standard output formats are incompatible combining a macOS-based system with a Linux-based system, either one as source or target, is therefore not straightforward. There is a known clash between the output format of the GNU / Linux tool md5sum and the macOS / BSD tool md5. You should mind the general limitations of checksums, which are however not covered in this recipe. A common benchmark on a typical laptop is: 0.01 seconds per MB of data. Depending on your system’s resources (especially available CPU time), this calculation of checksums might take a while, however. The above assumes that you don’t have a problem with calculating the checksums sequentially. (for a resolution see section “Extendability of this recipe” below.) The above assumes that everything is placed in your home folder. This recipe in its current form has the following limitations: IMI EUBOPEN FAIR High-Content Screening data deposition IMI ReSOLUTE - transcriptomics datasetsĦ. IMI Oncotrack - clinical cohort datasetsĤ. IMI nd4bb - chemical activities datasetsģ. Moving to a semantically typed version - ISA in RDFĢ. Making a community compliant - ISA in JSON Improving dataset maturity - the MIAPPE use case Using OpenRefine and Karma for FAIRification
Clinical Genetic Information as FHIR JSONĥ. Extraction, transformation, and loading processġ4. An inventory of tools for converting your data to RDFġ2. Converting from proprietary to open formatġ1. Metadata profile for transcriptomicsĩ.4.2. Application ontology for metabolomicsĩ.4.1. Competency questions for the Ontology ROBOT use caseħ.11.2. Building an application ontology with ROBOTħ.11.1. Introduction to terminologies and ontologiesĤ. Interlinking data from different sourcesģ. Depositing in Zenodo generic repositoryĨ.9.2. InChI and SMILES identifiers for chemical structuresĤ. Practical Considerations for a CRO to do FAIRĢ. Public Knowledge Graphs for Life Sciences
